What we know about nonsteroidal anti-inflammatory drug hypersensitivity
Article information
Abstract
Nonsteroidal anti-inf lammatory drugs (NSAIDs) are widely prescribed for the treatment of inflammatory diseases, but their use is frequently related to hypersensitivity reactions. This review outlines our current knowledge of NSAID hypersensitivity (NHS) with regard to its pathogenic, molecular, and genetic mechanisms, as well as diagnosis and treatment. The presentation of NHS varies from a local (skin and/or airways) reaction to systemic reactions, including anaphylaxis. At the molecular level, NHS reactions can be classified as cross-reactive (mediated by cyclooxygenase inhibition) or selective (specific activation of immunoglobulin E antibodies or T cells). Genetic polymorphisms and epigenetic factors have been shown to be closely associated with NHS, and may be useful as predictive markers. To diagnose NHS, inhalation or oral challenge tests are applied, with the exclusion of any cross-reactive NSAIDs. For patients diagnosed with NHS, absolute avoidance of NSAIDs/aspirin is essential, and pharmacological treatment, including biologics, is often used to control their respiratory and cutaneous symptoms. Finally, desensitization is recommended only for selected patients with NHS. However, further research is required to develop new diagnostic methods and more effective treatments against NHS.
INTRODUCTION
Nonsteroidal anti-inflammatory drugs (NSAIDs) are widely prescribed for the treatment of various inflammatory diseases. However, NSAIDs can cause drug hypersensitivity reactions, varying from local reactions in the skin and/or airways to systemic symptoms, including life-threatening anaphylaxis [1,2]. NSAID hypersensitivity (NHS) has been reported in 4.3% to 11% of patients with asthma [3] and in 27% to 35% of patients with chronic urticaria (CU) [4]. Moreover, NHS contributes to 13.3% of anaphylaxis cases in Korean adults [5]. While various terms (e.g., sensitivity, idiosyncrasy, or intolerance) have been used to describe unintended and unpredictable adverse drug reactions, the term “hypersensitivity” seems appropriate to describe the adverse reaction to NSAIDs, as its pathogenesis can involve both immunological and nonimmunological reactions [6,7].
NHS reactions may be classified as cross-reactive against several and chemically unrelated NSAIDs, or selective for a single compound or a group of chemically related compounds. Cross-reactive NHS reactions can manifest as three major clinical phenotypes: (1) NSAID-exacerbated respiratory disease (NERD), occurring in patients with underlying chronic upper and lower airway respiratory diseases; (2) NSAID-exacerbated cutaneous disease (NECD), occurring in patients with a history of CU; and (3) NSAID-induced urticaria/angioedema (NIUA), occurring in otherwise healthy subjects (without a history of CU) [2,8]. Anaphylaxis is commonly seen in NIUA. In addition, cross-reactive NHS may induce both respiratory and cutaneous symptoms in a subgroup of patients [9]. Selective NHS reactions manifest as two clinical phenotypes: (1) single NIUA/anaphylaxis reactions, which develop immediately after drug intake (immunoglobulin E [IgE]-mediated); and (2) NSAID-induced delayed hypersensitivity reactions, which develop more than 24 hours after drug intake (T cell-mediated) [2].
The pathogenesis of NHS reactions is thought to be related to various mechanisms, including: (1) a non-immunological mechanism, through the inhibition of enzymes involved in the arachidonic acid (AA) pathway; and (2) an immunological mechanism, involving the production of drug-specific IgE antibodies or T cell activation. Moreover, genetic polymorphisms and epigenetic factors have been implicated in the pathogenesis of, and susceptibility to, NHS. This review summarizes current knowledge regarding the pathogenic mechanisms, associated genetic and epigenetic factors, diagnostic methods, and treatments for patients with NHS.
PATHOGENIC MECHANISMS OF NHS
The proposed pathogenic mechanisms and clinical manifestations of NHS are shown in Table 1 and Fig. 1 [10]. NHS reactions can be subdivided as cross-reactive, in which the cyclooxygenase 1 (COX-1) pathway is inhibited, or selective, resulting from an immunological (allergic) reaction to the drugs and characterized by IgE-mediated (acute) or T cell-mediated (delayed) reactions [2,11-13]. In addition, novel mechanisms related to oxidative stress and platelet activation that could contribute to the pathogenesis of NHS have been reported [14-17].
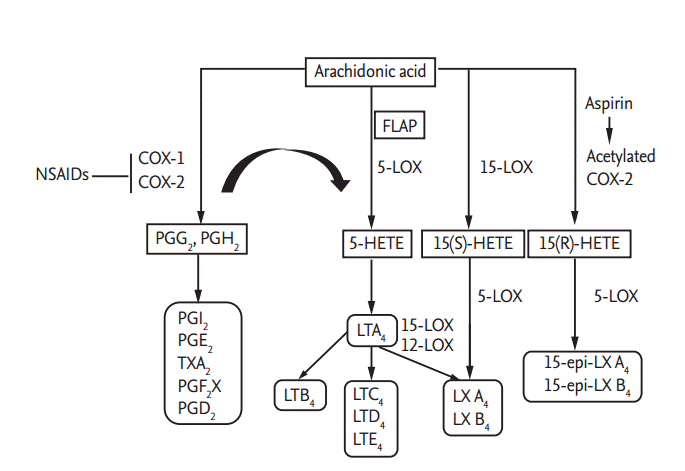
Arachidonic acid (AA) metabolism involved in the pathogenic mechanisms of nonsteroidal anti-inflammatory drug (NSAID) hypersensitivity. AA can be metabolized by either the cyclooxygenase (COX) pathway or 5-lipoxygenase (5-LOX) pathway [25]. Inhibition of COX enzymes by NSAIDs shifts the conversion of AA to the 5-LOX pathway, leading to decreased production of prostaglandins (PGs) and increased production of cysteinyl leukotrienes (LTs). Lipoxin (LX) A4/B4 are synthesized either from LTA4 or from 15S-hydroxyeicosatetraenoic acid (15(S)-HETE). In addition, aspirin acetylates COX-2, transforming AA to 15(R)-HETE and then to 15-epi-LX by 5-LOX [125]. In NSAID-exacerbated respiratory disease patients, the production of LX and 15-epi-LX, which have anti-inflammatory effects, could be decreased, possibly due to the excessive usage of AA for LT synthesis. FLAP, 5-lipoxygenase-activating protein.
Metabolism of arachidonic acid and the effects of NSAIDs
AA can be metabolized by either the COX pathway or the 5-lipoxygenase (5-LOX) pathway, with conversion to prostanoids or cysteinyl leukotrienes (CysLTs), respectively [18]. There are two isoforms of COX: COX-1 (expressed constitutively in most cells) and COX-2 (expressed either constitutively or in responses to inflammatory stimuli) [2]. The COX enzymes catalyze AA to prostaglandin (PG) G2 and then to PGH2, proceeding to the syntheses of PGE2, PGI2, PGD2, PGF2α, and thromboxane A2 (TXA2) by tissue-specific isomerase and synthase enzymes [18]. During prostanoid syntheses, COX-1 couples preferentially to thromboxane synthase (TBXAS), PGF synthase, and cytosolic PGE synthase (PGES) isozymes, while COX-2 shows preferential coupling with PGI synthase and microsomal PGES isozymes [19]. In the 5-LOX pathway, 5-LOX converts AA into LTA4, which is then converted to LTB4 (by LTA4 hydrolase) or catalyzed to LTC4 (by LTC4 synthase), followed by LTD4 and then E4 syntheses [20].
NSAIDs act by suppressing the activities of (1) COX isoenzymes, and (2) the enzymes responsible for prostanoid biosynthesis from AA [18,21,22]. Certain NSAIDs preferentially inhibit COX-1 and only partially inhibit COX-2 (e.g., aspirin, indomethacin, naproxen, and diclofenac); thereby, inhibiting production of protective PGs. Newer NSAIDs that inhibit COX-2 primarily (e.g., nimesulide, meloxicam) or specifically (e.g., celecoxib, rofecoxib) can suppress the inflammatory prostanoids, and only slightly decrease protective PG production [2].
COX-1 inhibition
By suppressing COX-1 activity, aspirin and some classical NSAIDs inhibit the synthesis of PGs, especially PGE2; thereby, increasing the production of CysLTs (LTA4, LTB4, LTC4, LTD4) [2,17,23,24]. PGEE2 has a protective effect against bronchoconstriction, releases inflammatory mediators from mast cells, and recruits immune cells to inflammatory sites [25,26]. On the other hand, CysLTs induce bronchoconstriction, airway inflammation, cell recruitment, and platelet activation [27-30]. In addition, aspirin can acetylate COX-2, leading to the generation of 15-hydroeicosatetraenoic acid (15-HETE), which is then transformed into lipoxin (LX) or 15-epimer-LX (15-epiLX) by 5-LOX. Both LX and 15-epi-LX exert anti-inflammatory effects by inhibiting neutrophil and eosinophil chemotaxis, as well as blocking LTC4-induced bronchoconstriction [23,31-33]. However, in NERD patients, the levels of LXA4 (downstream product of LXs) and 15-epi-LX in plasma and nasal lavage fluid were reported to decrease after lysine-aspirin nasal challenge [23,31]. A possible reason for the reduced LXA4 and 15-epi-LX levels in these patients is that the cellular AA resources are already overused for the synthesis of CysLTs [23].
IgE-mediated allergic reactions
NSAIDs can also elicit the production of specific IgE antibodies, which bind to their high-affinity receptors (FceRIα and FcεRII) on the surface of mast cells and basophils and provide multivalent binding sites for drug antigens. When the drug antigen is cross-linked with IgE bound to the receptors, the mast cells or basophils are stimulated to release preformed mediators (e.g., histamine) and produce new mediators [34]. Although a few studies showed high serum-specific IgE level production by NSAIDs, these results were not replicated and require further investigation [13,35-37].
Despite the conflicting results, one serum-specific IgE that has been confirmed to be involved in NHS is the IgE against thyroid peroxide (TPO), which was first discovered by Altrichter et al. [38] in 2011. More recently, high TPO-specific IgE levels were found in patients with NIUA and NECD compared to healthy controls [39]. Moreover, when TPO was added to basophils isolated from these NIUA and NECD patients, the expression of CD203c (a surface marker for basophil activation) increased [39]. These findings suggest a role of TPO-specific IgE in the pathogenesis of NECD/NIUA via activation of basophils or mast cells [39]. Furthermore, crosslinking of an allergen to the neighboring IgE molecules bound to FceRI can trigger calcium influx, resulting in basophil degranulation [40]. Moreover, in patients with CU and food-dependent, exercise-induced anaphylaxis, aspirin treatment was found to increase phosphorylation of the spleen tyrosine kinase (Syk) in IgE-sensitized basophils [16]. Finally, other NSAIDs (e.g., diclofenac, salicylate, and ketoprofen) were also found to indirectly enhance IgE-mediated histamine release from human basophils through activation of the Syk kinase pathway [40,41].
T cell-mediated mechanisms
NSAIDs can induce delayed-type reactions mediated by T cell activation [2,42]. In the context of maturation signals resulting from drug-related stress, disease, or trauma, dendritic cells (DCs) in the skin and mucous membrane recognize and transport drug antigen complexes to the regional lymph nodes [43]. In the lymph nodes, DCs introduce the drug antigens to naïve T lymphocytes and stimulate the production of antigen-specific T cells. Subsequently, drug antigen-specific T cells migrate to the target organs and secrete cytokines and cytotoxins upon re-exposure to the drugs [43]. Based on the main effector cells involved, T cell-mediated NHS mechanisms are classified into four subtypes: (1) IVa, involving type 1 helper (Th1) cells and monocytes and the production of interferon g, interleukin 1 (IL-1), and IL-2 cytokines; (2) IVb, involving Th2 cells and eosinophils and the production of IL-4, IL-5, and IL-13; (3) IVc, involving cytotoxic T cells and the production of perforin, granzyme B, and Fas ligand; and (4) IVd, involving CD4+ T cells, CD8+ T cells, and neutrophils, and the production of IL-8 and granulocyte-macrophage colony-stimulating factor [7,42].
Oxidative stress and apoptosis in NHS
Another mechanism underlying NHS involves oxidative stress and the induction of pro-inflammatory mediator production (cytokines and chemokines), which can aggravate airway inflammation, bronchospasm, and mucin secretion [14]. For example, Antczak et al. [44] showed that the levels of 8-isoprostanes, metabolized products from AA that are induced by reactive oxygen species, are increased in the exhaled breath condensate of patients with NERD [45], suggesting a role of oxidative stress in NHS. Moreover, aspirin can trigger apoptosis by inducing oxidative stress as well as by inhibiting expression of the anti-apoptotic protein, Beclin-2 (Bcl2) [17]. NSAIDs may reduce Bcl2 expression by blocking the IL6-IL6R-STAT3 signaling pathway. In turn, decreased Bcl2 expression initiates tumor necrosis factor (TNF)-related apoptosis-inducing ligand and TNF-a generation, which then elicit apoptosis [17].
Platelet activation in NHS
While platelet activation has long been known to be involved in the pathogenic mechanisms of asthma and urticaria [46,47], its role in NHS has not been studied in depth. However, Palikhe et al. [15] recently found a higher level of soluble P-selectin, which is a marker for activated platelets, in patients with NECD than in those with aspirin-tolerant CU or healthy controls. In addition, while aspirin treatment abolished the expression of soluble P-selectin and P2Y12 on platelets in aspirin-tolerant CU patients, no such inhibitory effect was observed in platelets from NECD patients [15]. These findings suggest that platelet activation contributes to the pathogenesis of NECD.
GENETIC AND EPIGENETIC FACTORS INVOLVED IN NHS
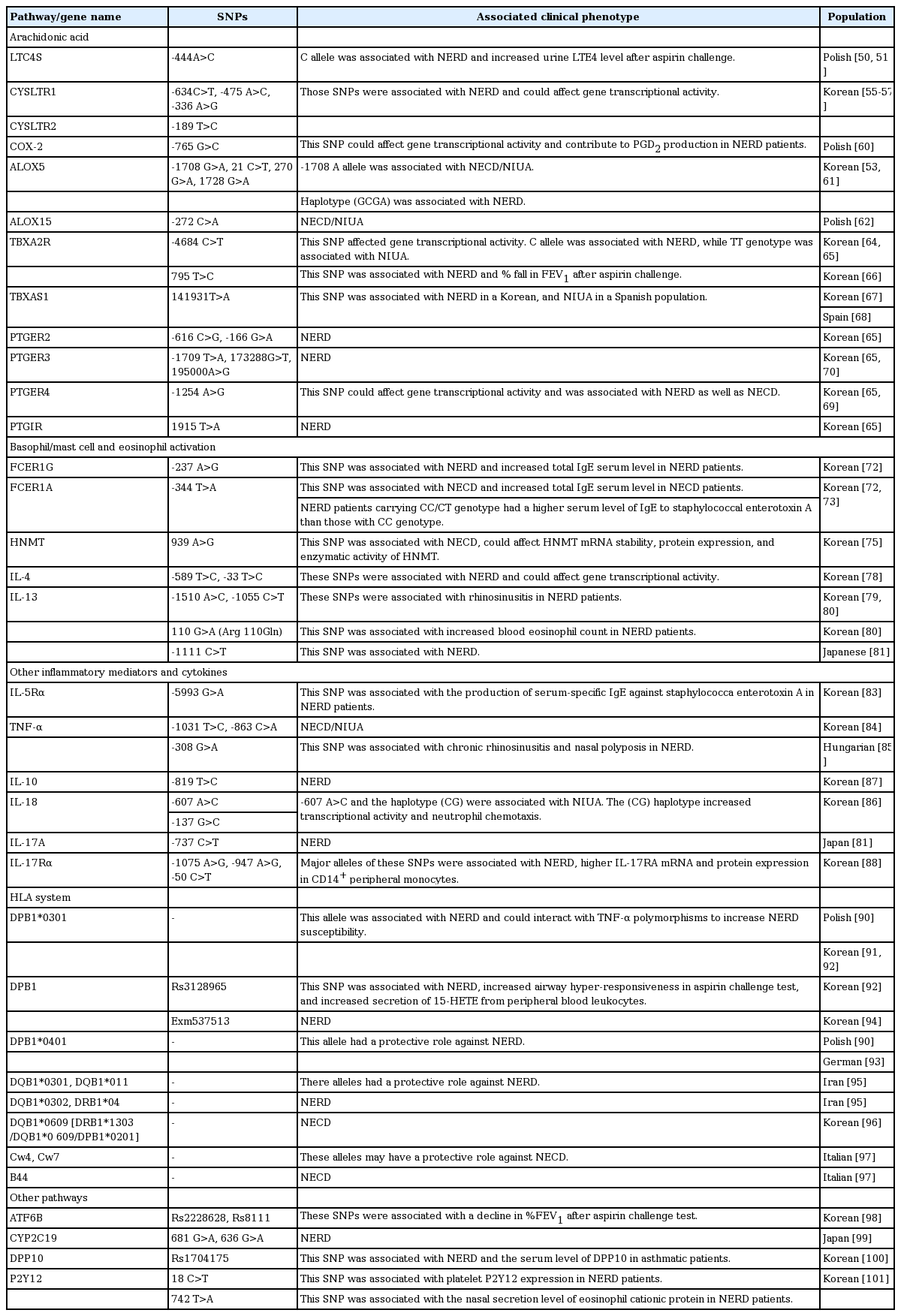
A family history of aspirin intolerance has been reported in 6% of patients, thereby suggesting a role of genetic factors in NHS pathogenesis [48]. However, most genome-wide association and case-control studies have focused on NERD, and only a limited number of studies have investigated genetic factors involved in NECD/NIUA. These studies concentrated on the genes related to the suspected pathomechanisms of the disease, including AA metabolism pathways, IgE, inflammatory mediators, and cytokines, the human leukocyte antigen (HLA) system, and other miscellaneous factors [8,49]. Genetic polymorphisms that have been found to be associated with NHS are shown in Table 2. In addition, epigenetic factors have recently been found to be associated with the pathogenic mechanisms of NHS, which are also summarized in this section.
Genes involved in AA metabolism pathways
Early studies on NHS-associated genes focused on the gene encoding leukotriene C4 synthase (LTC4S). The C allele of LTC4S -444A>C (rs730012) was reported to be a risk factor for NERD, and was associated with an increase in urine LTE4 level after aspirin challenge test [50]. In addition, NIUA was found to aggregate in Polish families inheriting the LTC4S -444C allele [51]. However, these findings were not replicated in American, Japanese, or Korean populations [52-54]. Instead, an association of NERD with three single nucleotide polymorphisms (SNPs) in the CysLT receptor 1 gene (CYSLTR1 -634C>T, -475A>C, and -336A>G) and one SNP in CYSLTR2 (-189T>C) was identified in Korean populations. These SNPs were also found to affect gene transcriptional activity [55-57], consistent with the observed increases in CysLTR1 and CysLTR2 expression found in nasal mucosal inflammatory cells of NERD patients [58,59].
In Polish subjects with NERD, a promoter SNP in COX-2 (-765G>C) was found to affect transcriptional activity and contribute to the production of PGD2 [60], while four SNPs in the 5-LOX gene (ALOX5 -1708G>A, 21C>T, 270G>A, and 1728G>A) were associated with NERD in a Korean population [53]. The ALOX5 -1708A allele was also associated with NECD/NIUA in a Korean population [61]. In addition, a promoter SNP in ALOX15 (-272C>A) was found to be associated with NECD/NIUA in a Spanish population. However, associations of ALOX12 and ALOX15 polymorphisms with NHS were not observed in a Korean population [62,63].
The transcriptional function of the -4684C>T SNP in the TXA2 receptor gene (TBXA2R) was reported previously, and was associated with NIUA [64]. Consistently, the TBXA2R -4684C allele was shown to be associated with NERD, while the TBXA2R -4684TT genotype was associated with NIUA in a Korean population [64,65]. A nonsynonymous SNP of TBXA2R (795T>C) was also found to be associated with NERD, as well as with a drop in forced expiratory volume in 1 second (FEV1) after inhaled aspirin challenge [66]. Similarly, the SNP in intron 9 (rs6962291, 141931T>A) in the TBXA1 synthase gene (TBXAS1) was shown to be associated with NERD in a Korean population [67], as well as with NIUA in a Spanish population [68]. In a Korean population, NERD was significantly associated with genetic polymorphisms of the prostanoid receptors, prostaglandin E receptor 2 (PTGER2) (-616C>G, -166G>A), PTGER3 (-1709T>A), PTGER4 (-1254A>G), and PTGIR (1915T>A) [65]. Moreover, PTGER4 -1254A>G was found to affect transcriptional activity and was associated with NECD [69]. Another study indicated an association of NERD with rs7543182 (173288G>T) and rs959 (195000A>G) in PTGER3 [70].
Genes involved in mast cell/basophil and eosinophil activation
The high-affinity IgE receptor (FcεRI) plays a crucial role in the activation and degranulation of mast cells and basophils, resulting in the release of a variety of cytokines and leukotrienes [71]. Palikhe et al. [72] demonstrated an association of FCER1G -237A>G with NERD and increased total IgE serum levels in a Korean population with NERD. On the other hand, the C allele of FCER1A -344T>C was associated with NECD and increased total serum IgE level as well as atopy rate in NECD patients [73]. Although no association of FCER1A -344T>C with NERD was found, the FCER1A -344CT/TT genotype was associated with a higher serum level of IgE specific antibody to Staphylococcal enterotoxin A compared to the CC genotype [72]. Nevertheless, these findings could not be replicated in a recent study with a small population of Taiwanese CU patients [34].
The cellular level of histamine, a key mediator released from basophils/mast cells, is regulated by histamine N-methyltransferase (HNMT) [74]. Therefore, the HNMT 939A>G polymorphism may contribute to NECD by influencing HNMT mRNA stability, protein expression, and HNMT enzymatic activity [75,76]. In addition, IL-4 and IL-13 are important cytokines involved in mast cell, basophil, and/or eosinophil activation and function, and share a common receptor subunit (IL-4Rα) [77]. The IL-4 -589T>C and -33T>C polymorphisms could affect the promoter transcriptional activity, and were shown to be associated with NERD risk [78]. Although associations of IL-13polymorphisms (-1510A>C, -1055C>T, and 110G>A [Arg110Gln]) with NERD and NECD were not observed in a Korean population, higher frequencies of rhinosinusitis were noted in NERD patients carrying the IL-13 -1510A allele, as well as the -1055CC genotype, compared to non-carriers [79,80]. Moreover, the GG genotype of IL-13 110G>A was significantly associated with an increased blood eosinophil count in NERD patients [80]. In addition, in a Japanese population, the minor allele of IL-13 -1111C>T was reported to be associated with NERD [81]. Finally, IL-5 and its receptor IL-5R are necessary for the survival and activation of eosinophils [82], and Losol et al. [83] recently described an association between an IL-5Rα polymorphism (-5993G>A) and the production of serum-specific IgE antibodies against Staphylococcal enterotoxin A. However, no association of this IL-5Rα polymorphism with NERD susceptibility was found.
Other inflammatory mediators and cytokine-related genes
Among the five SNPs of the TNF-α gene investigated in our previous study (TNF-α -1031T>C, -863C>A, -857C>T, -308G>A, and -238G>A), only two (TNF-α -1031T>C and -863C>A) were significantly associated with NECD/NIUA risk [84]. Associations of chronic rhinosinusitis (CRS) and nasal polyps with TNF-α -308G>A in NERD patients were reported in a Hungarian population, although no association with NERD susceptibility was observed [85]. Kim et al. [86] demonstrated an association of NIUA with IL-18 -607A>C, as well as with the haplotype (CG) of IL-18 -607A>C and -137G>C, which could increase transcriptional activity and neutrophil chemotaxis. In addition, associations of NERD with other cytokine gene polymorphisms, including IL-10 (-819T>C), IL-17A (-737C>T), and IL-17Rα (-1075A>G, -947A>G, -50C>T) were also reported in some studies [81,87,88]. These findings suggest that NHS is associated with numerous genetic polymorphisms of inflammatory cytokines, the functions of which in the pathogenesis of NHS should be investigated further.
HLA genes
HLA molecules play a crucial role in T cell activation. Classical HLA class I molecules (HLA-A, HLA-B, and HLA-C) are responsible for presenting drug antigens to CD8+ T cells, while HLA class II molecules (HLA-DP, HLA-DQ, and HLA-DR) present antigens to CD4+ T cells [89]. Among them, HLA class II alleles are well-known genetic factors associated with NHS. The association of NERD with HLA-DPB1*0301 was first reported in a Polish population, and these findings were then replicated in a Korean population [90,91]. Interestingly, HLA-DPB1*0301 combined with certain TNF-α polymorphisms increases the susceptibility to NERD [91]. More recently, the A allele of rs3128965 in HLA-DPB1 was found to be associated with NERD, increased airway hyper-responsiveness after inhaled aspirin challenge test, and increased secretion of 15-HETE from peripheral blood leukocytes treated with AA [92]. In contrast, the HLA-DPB1*0401 allele was reported as a protective factor against NERD susceptibility in studies with Polish and German populations [90,93]. A novel exonic polymorphism (exm537513) in HLA-DPB1 associated with NERD was also recently identified [94]. In addition, the combination of exm537513 with six other exonic SNPs from other genes (exm83523, exm1884673, exm538564, exm2264237, exm396794, and exm791954) provides good predictive value for NERD (area under the curve of 0.75 with 34% sensitivity and 93% specificity) [94]. In addition, Esmaeilzadeh et al. [95] reported that HLA-DQB1*0302, HLA-DRB1*04, and their haplotype were risk factors of NERD, while HLA-DQB1*0301 and HLA-DRB1*011 were negatively associated with NERD.
NECD has also been found to be associated with HLA-DRB1*1302, HLA-DQB1*0609, and the haplotype of HLA-DRB1*1303/DQB1*0609/DPB1*0201 in a Korean population [96]. Furthermore, NECD has also been reported to be associated with HLA class I alleles. Pacor et al. [97] showed that HLA-Cw4 and HLA-Cw7 could have a protective role against NECD, while HLA-B44 was a risk factor. The study by Pacor et al. [97] indicates that HLA class I molecules are involved in the pathogenesis of NHS, and further investigations are required.
Genes related to other pathways
Several genetic risk factors for NERD related to other mechanisms have also been identified. For example, polymorphisms of the activating transcription factor 6β gene (ATF6B rs2228628 and rs8111) were associated with decline in FEV1 % after inhaled aspirin challenge test [98]. Moreover, two SNPs in cytochrome P450 CYP2C19 (681G>A and 636G>A) were found to be associated with NERD and the predicted FEV1 % value [99]. Kim et al. [100] recently reported an association of dipeptidyl-peptidase 10 gene (DPP10) rs1704175 (TT genotype) with NERD, and the TT genotype was associated with a higher serum DPP10 level compared to the CC/CT genotypes in asthmatic patients. The purinergic receptor gene polymorphism (P2RY12 742T>C) was associated with the nasal secretion level of an eosinophil cationic protein, while P2RY12 18C>T was associated with platelet P2Y12 expression level in NERD patients [101]. Nevertheless, the underlying mechanisms by which these genes contribute to NERD pathogenesis remain to be elucidated.
Epigenetic mechanisms
A recent study evaluated DNA methylation levels in nasal polyp tissues of patients with NERD and aspirin-tolerant asthma (ATA) [102]. Comparison of the NERD and ATA groups indicated increased methylation levels of 332 loci in 296 genes, while 158 loci in 141 genes had decreased methylation levels. In addition, four genes involved in AA metabolism (PGD synthase [PGDS], PTGES, ALOX5 activating protein [ALOX5AP], and leukotriene B4 receptor [LTB4R]) were found to have different DNA methylation levels between NERD and ATA patients. These findings highlight the value of nasal polyp profiles for suggesting potential susceptibility genes and pathways involved in the pathogenesis of NERD [102].
MANAGEMENT OF PATIENTS WITH NHS
Clinical manifestations
Symptoms of acute NHS include rhinitis/asthma, urticaria/angioedema, and anaphylaxis [82,103]. Patients with NERD have hypersensitivity in the upper (CRS, nasal polyps) and lower (asthma) airways, called Samter’s triad [104]. In cases of acute NERD, rhinorrhea and nasal congestion followed by dyspnea and chest tightness appear within 30 to 120 minutes after administration of NSAIDs. The chronic form of NERD, i.e., preexisting inflammation of the upper and lower airways that is exacerbated by ingestion of NSAIDs, usually appears in the third decade of life, and is characterized by a variety of symptoms, including nasal congestion, hyposmia, pansinusitis, laryngospasm, and bronchospasm [105,106]. The phenotype of the disease progresses to severe and corticosteroid-dependent asthma with NHS, and NERD can persist throughout life [107,108]. Cutaneous (such as urticaria) and gastric symptoms may accompany the respiratory symptoms in these patients.
In NECD, symptoms occur within 1 to 4 hours after ingestion of a culprit drug, whereas delayed reactions may occur up to 24 hours. Hives usually disappear within several hours but may persist for days, and some NECD patients may concurrently develop respiratory symptoms [4]. In 30% of CU patients, hypersensitivity reactions occur following ingestion of aspirin or NSAIDs [109]. While aggravation of underlying urticaria by ingestion of the culprit drug can occur in these CU patients, the symptoms may fluctuate depending on how well the underlying disease is controlled [110].
Anaphylaxis is another type of acute reaction in NHS, which is commonly combined with NIUA. Within a few minutes of ingestion or administration of the culprit drug, generalized urticaria and skin swelling may progress into anaphylaxis. Some patients develop both symptoms of NERD and CU [24]. Drugs notorious for causing anaphylaxis are pyrazolones, ibuprofen, diclofenac, aspirin, and paracetamol [24]. In delayed reactions, symptoms occur within days or even weeks after exposure, but may be induced earlier by reintroduction. Cutaneous reactions (such as fixed drug eruption, severe bullous cutaneous reaction, and maculopapular drug eruption) mostly occur, although rarely organ-specific symptoms (aseptic meningitis, pneumonitis, or nephritis) are reported [24,111].
Diagnosis
The initial step in the diagnosis of NHS is examination of clinical history. Carefully obtained information on symptom descriptions, symptom chronology (including order of incidence or previous exposure to the culprit drugs), and underlying comorbidities (including asthma, CRS, nasal polyps, or CU) are key for NHS diagnosis [6,112]. Adverse reactions to NSAIDs are those where the patient had an adverse drug reaction within 6 hours of NSAID intake, or if such a reaction was recorded by a physician, or if a reaction reoccurred to the same or a distinct NSAID at different times [109]. However, further diagnostic modalities should be applied, as clinical history is never enough to identify the culprit drugs or mechanisms of the reaction. Therefore, skin prick and intradermal tests are useful tools to confirm or exclude sensitization in acute reactions related to an IgE-dependent mechanism, while a patch test is used to identify a delayed reaction related to T cell-mediated responses [6]. However, the sensitivity and specificity of these tests remain unclear and, to date, no standardized tests have been established [6,24]. Moreover, as an in vitro study, the basophil activation test may provide useful information; however, it is not used widely due to its low sensitivity and difficulties in setting up the method.
While skin or in vitro tests are useful for confirming the culprit drugs, the oral challenge test remains the gold standard for NHS diagnosis [113]. Indeed, an l-lysine-aspirin challenge test and/or aspirin oral challenge test should be performed to confirm the diagnosis of potential NERD patients [106,113]. These oral challenge tests to NSAIDs show very high sensitivity and specificity, exceeding 90% [2]. For the aspirin oral challenge test, the European Academy of Allergy and Clinical Immunology/Global Allergy and Asthma European Network (EAACI/GA2 LEN) guidelines outline a 2-day protocol where four placebo capsules are administered on the first day, and then four increasing doses of aspirin (71, 117, 312, and 500 mg) are given within a 2-hour interval on the second day, followed by FEV1 evaluation every 30 minutes [113]. In Korea, the l-lysine-aspirin bronchoprovocation test is widely applied to confirm a diagnosis of NERD; however, to perform these bronchoprovocation tests, the patient should be in a stable condition and the tests should not be performed in a patient with FEV1 < 70%. Moreover, the bronchoprovocation test should be implemented under the guidance of physicians and technicians equipped with emergency resuscitative equipment [113,114]. The contraindications of challenge tests are acute severe exacerbations, a history of anaphylaxis, or severe cutaneous adverse reactions. It is important to note that although negative results to challenge test may be observed in highly suspicious cases, the diagnosis of NHS should not be excluded, especially in patients on long-term corticosteroid treatment or those with well-controlled underlying diseases.
Recently, Kowalski et al. [2] outlined a simplified, practical diagnostic algorithm for NHS, as follows: (1) the allergist should evaluate if the symptoms are predictable (intolerant) or unpredictable (hypersensitivity); (2) the onset time of the reactions should be assessed; and (3) the pattern of clinical symptoms and underlying chronic disease should be analyzed. In addition, identifying a history of tolerance or intolerance to other NSAIDs helps suggest appropriate NSAIDs for use in challenge tests to confirm or exclude cross-reactivity with other NSAIDs.
Treatment
Immediate NSAID cessation and strict avoidance of the culprit drug and any cross-reacting NSAIDs are of the utmost importance in both early and delayed NHS reactions. Moreover, written information, including lists of culprit and alternative medications, should be provided to NHS patients [7,112].
Most NERD patients have moderate to severe asthma, and medium to high doses of inhaled corticosteroid/long-acting β agonist and leukotriene modifiers should be maintained to control lower respiratory symptoms. The upper airway symptoms related to CRS and/or nasal polyps should be controlled to improve bronchial symptoms, either medically using intranasal corticosteroids, or surgically if necessary [115]. In cases of severe asthma with frequent asthma exacerbations, biologics, such as anti-IgE antibodies, may have beneficial effects for controlling upper and lower respiratory symptoms [116-118]. The suggested biologics for use in these patients include omalizumab, mepolizumab, and benralizumab [119-121]. For the treatment of CRS/nasal polyps, along with medical treatment (e.g., intranasal corticosteroids), consultation with an ear, nose, and throat specialist for surgical treatment, such as sinus endoscopy, is advised [122].
Systemic steroids and antihistamines may be used during acute, severe cutaneous reactions. However, in severe and delayed reactions (e.g., Stevens-Johnson syndrome [SJS] and toxic epidermal necrolysis [TEN]) withdrawal of the culprit drugs is the only proven treatment to decrease fatality. Although other treatments have been suggested for severe adverse reactions (including systemic corticosteroids, intravenous Igs, or immunosuppressive drugs [e.g., cyclosporin and infliximab]), these treatments remain controversial [24].
Finally, desensitization, which involves the induction of a temporary state of tolerance/unresponsiveness to the drug (mostly aspirin) responsible for NHS, may be an optional treatment for NERD and is expected to improve upper and lower respiratory symptoms [123,124]. Once desensitized, patients should take the drug regularly to maintain tolerance, as intolerance could recur after 2 to 5 days of cessation [124]. Oh et al. [125] reported two cases of NECD that were successfully controlled by aspirin desensitization. Unfortunately, protocols for drug desensitization have not been standardized to date; however, slower protocols tend to be more effective than rush protocols [114]. Despite these successes, the EAACI guidelines indicate that desensitization is absolutely contraindicated in cases of severe or threatening delayed drug hypersensitivity reactions (such as SJS, TEN, or drug reactions with eosinophilia and systemic symptoms) [114].
CONCLUSIONS
NHS is a heterogeneous disease with several clinical phenotypes involving various pathogenic mechanisms. In addition to COX pathway alterations, NHS is related to immune responses (including IgE production and T cell activation), oxidative stress, and platelet activation. While recent genetic studies have identified some predictive markers for NHS susceptibility and treatment response in different populations, these require further investigation. In future, an improved understanding of the functional and genetic/epigenetic pathogenic mechanisms of NHS will help in the development of new diagnostic methods and effective treatments.
Notes
No potential conflict of interest relevant to this article was reported.
Acknowledgements
This study was supported by a grant from the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (H14C2628).