Relevance of vacA Genotypes of Helicobacter pylori to cagA Status and Its Clinical Outcome
Article information
Abstract
Background :
Determination of vacA mosaicism may be important because specific Helicobacter pylori vacA genotype can be used to predict different clinical outcome. The aim of this study was to assess the relationship of vacA genotypes of Helicobacter pylori to cagA status and its development of peptic ulcer diseases in Korean patients.
Methods :
Gastric biopsy specimens were obtained from 53 patients with gastric ulcer (GU), 57 with duodenal ulcer (DU) and 26 with chronic gastritis (CG) patients; all patients were infected with Helicobacter pylori. Bacterial mRNAs in the gastric mucosa were amplified by RT-PCR, using synthetic oligonucleotide primers specific for the vacA and the cagA gene. Patients with vacA s1 subtype were further examined to determine whether they had s1a or s1b subtype.
Results :
There was no correlation in frequency of vacA s1 and/or s1a genotype between CG and either GU or DU, as the vacA s1 and s1a/m1 were present in the majority of strains independent of clinical status (s1 ; 100.0% versus 94.3 % or 93.0 % and s1a/m1 : 76.9% versus 62.3% or 64.9%, respectively). Likewise, there was no difference in the prevalence of the cagA gene between CG and either GU or DU patients (92.3% versus 90.6% or 98.2%, respectively). In addition, the cagA-negative status did not predict the presence of vacA s2 genotype.
Conclusion :
These results strongly suggest that either cagA or vacA s1 and/or s1a is not proved to be a useful marker to distinguish disease-specific Helicobacter pylori strains for the development of peptic ulcer diseases in Korean patients.
Introduction
Helicobacter pylori (H. pylori) infection may induce chronic gastritis, peptic ulcer diseases, gastric adenocarcinoma and gastric MALT lymphoma. However, it has not been solved why most subjects infected with H. pylori are not affected by peptic ulcer diseases or gastric carcinoma. The cause of these different clinical outcomes of H. pylori infection remains unclear and may include host and environmental factors1–3) and differences in the prevalence or expression of bacterial virulence factors. Infection with certain strains of H. pylori is related to more severe morbidity, whereas other variants appear less pathogenic4–9). Two characteristics previously linked to pathogenicity are production of a vacuolating cytotoxin and possession of cagA gene. Both of these characteristics have been known to be significant risk factors for developing more serious gastroduodenal diseases5,7,10–12). Recently however, there have been increasing reports that the cagA gene is not proved to be a useful marker to distinguish disease-specific H. pylori strains for the development of peptic ulcer diseases 13–16).
Approximately 50% of clinical isolates of H. pylori produce VacA protein that causes vacuolar degeneration in several mammalian cell lines in culture, while all strains have the gene encoding the vacuolating cytotoxin, VacA17). Infection by a subset of H. pylori strains, referred to as type I, which contain a pathogenicity island in the chromosome and which are strongly toxigenic, can cause more severe pathology than type II, non-toxigenic strains12). Furthermore, intragastric administration of highly purified VacA caused epithelial erosions and ulcerations similar to those seen in mice infected with type I H. pylori, strains18). The genetic sequence of the vacA varies markedly between type I and type II strains of H. pylori especially in the region encoding the signal sequence and the mid-region of the gene18). This mosaic pattern of vacA alleles is similar or dissimilar between bacterial strains. All combinations of the three signal sequence types (s1a, s1b, or s2) and two mid-region types (m1 or m2) are found, implying horizontal acquisition of genetic elements and recombination between H. pylori strains in vivo18). Classification of vacA mosaicism may be important because specific H. pylori vacA genotypes can be used to predict different clinical status of H. pylori infection. Atherton et al.8,19) reported that s1 and/or s1a genotype was more frequently associated with duodenal ulcer disease and also associated with more severe infiltration of neutrophils and lymphocytes in human gastric antral mucosa than s1b or s2 strains. They also found that cagA status was closely associated with the vacA s1 or the s2 allele. For example, 88% of USA strains with vacA s1 allele were found to be cagA-positive, and all 19 strains with vacA s2 allele were cagA-negative.
The present study aimed to determine the relevance of vacA genotypes of H. pylori to cagA status and its development of peptic ulcer diseases in Korean patients infected with H. pylori.
MATERIALS AND METHODS
1. Population Studied
The study population was made up of 136 patients (107 males and 29 females; mean age, 46.6 years; range 17 – 81). One hundred and ten patients were endoscopically and histologically confirmed to have benign gastric ulcer (GU, 53 patients) or duodenal ulcer (DU, 57 patients) in addition to being infected with H. pylori. The control group consisted of 26 patients with chronic gastritis (CG) who were also infected with H. pylori. Patients were excluded if they had a history of gastric surgery, active gastrointestinal bleeding or exposure to steroids or NSAIDs, H2-receptor antagonists, proton pump inhibitors or antimicrobial agents within 30 days prior to this study. Patients with any other chronic illness were also excluded.
2. Endoscopy and diagnosis of H. pylori infection
Gastroscopic examination was performed and the endoscopic findings were recorded. Biopsy specimens in each patient were obtained from gastric antrum within 2 cm proximal to the pylorus and from gastric midbody. Histological confirmation of H. pylori infection was done by the Warthin-Starry silver staining method. Individual mucosal specimen was evaluated by a single pathologist who was unaware of the PCR results. Microaerophilic culture under 37°C and rapid urease test were also performed in each patient. The remaining biopsy specimens from gastric antrum and midbody in each patient were frozen at −70°C for RNA extraction and RT-PCR to detect the presence of H. pylori genes. Patients were classified as H. pylori-positive when the culture was positive or if at least three of the four examinations, such as rapid urease test, histologic examination, RT-PCR examination for the presence of ureA and 16S-rRNA, gave positive results for H. pylori.
3. Detection of H. pylori cagA, vacA and s1a subtype expression
Detection of H. pylori ureA, cagA and 16S-rRNA of genomic DNA from clinical isolates and of cDNA from gastric mucosal biopsy specimens was performed by PCR and RT-PCR amplification by using oligonucleotide primers described in detail previously15). In brief, amplification of the cagA gene was done using the previously described primers F1 (5′-GATAACAGGCAAGCTTTTGAGG-3′) and B1 (5′-CTGCAAAAGATTGTTTGGCAGA-3′) with annealing at 55°C to amplify a 349-bp product from the middle of the cagA.
Twenty-eight histology-positive patients (10 GU, 10 DU, 8 CG) were selected to test cagA and vacA s1a allele both from clinical isolates and from the mucosa in order to confirm that the expression of each gene in the gastric mucosa reflected the genotype of the infecting organism. After confirming these results, RT-PCR amplifications were done for ureA, 16S-rRNA, cagA, and vacA genotypes in the remaining patients using mucosal biopsy specimens.
4. PCR-based vacA genotyping
Typing of the vacA signal sequence and mid-regions was performed by using oligonucleotide primer pairs based on published data17). The primer pairs VA1-F/VA1-R (5′-ATGGAAATACAACAAACACAC-3′ and 5′-CTGCTTGAATGCGCCAAAC-3′) and SS2-F/VA1-R (5′-GCTAACACGCCAAATGATCC-3′ and VA1-R as above) were used to distinguish between s1 and s2. Primer pairs VA3-F/VA3-R (5′-GGTCAAAATGCGGTCATGG-3′ and 5′-CCATGGTACCTGTAGAAAC-3′) and VA4-F/VA4-R (5′-GGAGCCCCAGGAAACATTG-3′ and 5′-CATAACTAGCGCCTTGCAC-3′) were used to amplify the ml and m2 mid-regions, respectively. Each strain was typed as s1a or s1b by performing two separate RT-PCRs, each using one primer specific to the signal sequence type SS1-F (5′-GTCAGCATCACACCGCAAC-3′), SS3-F (5′-AGCGCCATACCGCAAGAG-3′), and one conserved primer (VA1-R as above). Thermal cycling of each set of primers (0.5 M each) was performed at 95°C for 1 min, 52°C for 1 min, and 72°C for 1 min, for a total of 35 cycles.
5. Statistical Analysis
Pearson’s chi-square test was used to determine the significance of difference among three groups with a difference in p value less than 0.05 being considered significant.
RESULTS
Twenty-eight histology-positive cases which were also positive for the ureA gene, rapid urease test and culture were selected in order to confirm that RT-PCR detection of cagA and expression of vacA alleles in the gastric biopsy specimen reflected the genotype of the infecting organism. Complementary DNA was prepared from the corresponding gastric biopsy specimens from the same patient whose specimen was used as the template for PCR. With genomic DNAs, the cagA was positive in 26 (92.9%) and the vacA s1a was positive in 24 (82.8%) out of 28 isolates. Each of the 26 and the 24 corresponding gastric biopsy specimens was also positive for cagA and s1a by RT-PCR. Neither the cagA nor the vacA s1a gene was found in the remaining 2 and 4 corresponding gastric mucosa whose genomic DNAs did not possess either cagA or vacA s1a gene. These results suggest that cagA and vacA s1a of bacterial genomic DNA detected by PCR are identical with those from the gastric mucosal biopsy detected by RT-PCR. After confirming these results, RT-PCR amplification was done for ureA, 16S-rRNA, cagA gene and vacA signal sequence and mid-region alleles in the remaining patients using mucosal biopsy specimens.
1. vacA Genotyping
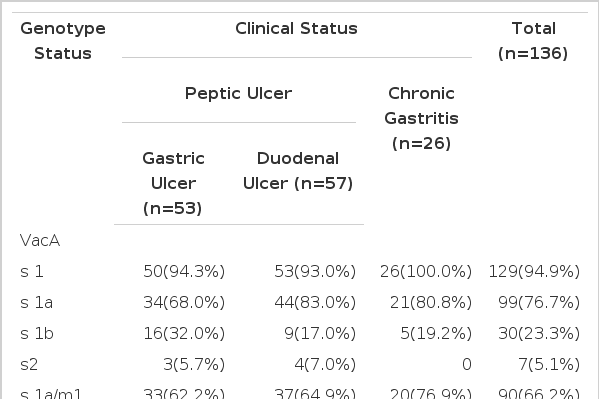
Signal sequence and mid-region genotypes of the vacA gene were obtained in all patients. One hundred and twenty nine (94.9%) out of 136 mucosal specimens had the s1 allele, of which 99 (76.7%) had the s1a subtype, whereas 30 (23.3%) had the s1b subtype. The remaining seven (5.1%) specimens had s2 subtype. Mid-region subtypes such as the vacA m1 and m2 were found in 118 (86.8%) and 18 (13.2%) out of 136 mucosal specimens, respectively. Overall, 90 (66.2%) strains were classified into the s1a/m1, 9 (6.6%) were s1a/m2, 28 (20.6%) were s1b/m1, 2 (1.5%) were s1b/m2, and 7 (5.1%) strains were s2/m2 (Table 1).
2. Association of vacA genotype with cagA gene
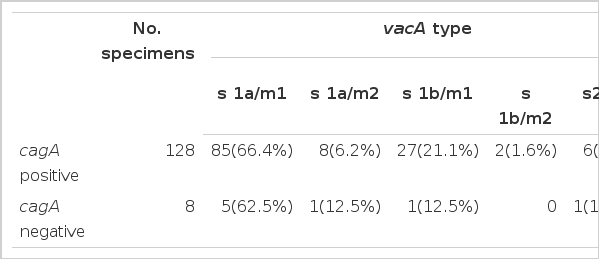
One hundred and twenty eight strains (94.1%) were positive for cagA gene by RT-PCR methods. Eight patients (5 GU patients, 1 DU, and 2 CG) were negative for cagA. The prevalence of cagA-positive was not different between patients with CG and patients with either GU or DU (92.3% vs. 90.6% and 98.2%, respectively). There was no correlation in the prevalence between the cagA and the vacA s1a, because vacA s1a genotype was found in 31 (64.6%) out of 48 GU patients, 43 (76.8%) out of 56 DU, and 20 (83.3%) out of 24 CG patients who were infected with cagA-positive H. pylori strains (Table 2 and 3). These results do not support the hypothesis that H. pylori strain possessing the cagA gene and the vacA s1a allele is a useful marker for the discrimination of specific clinical status. Likewise, the presence of cagA-negative status did not predict the presence of vacA s2 genotype because 7 out of 8 cagA-negative strains were the s1 (6 isolates with s1a and 1 with s1b) and 6 isolates with the s2 subtype were cagA-positive.
3. Association of vacA genotypes with the disease status
All of the CG patients had vacA s1 allele genotype. Twenty-one patients (80.8%) were positive for the s1a and 5 patients (19.2%) were positive for the s1b subtype. Fifty patients (94.3%) with GU had s1 allele, of which 34 (68.0%) had s1a and 16 (32.0%) had s1b subtype. In DU patients, 93.0% (53 out of 57 patients) had s1 genotype, consisting of 83.0% (44 out of 53) with s1a and 17.0%(9 out of 53) with s1b subtype.
DISCUSSION
Our study demonstrated that vacA gene was found in all H. pylori obtained from 136 mucosal specimens and all combinations of the signal sequence and the mid-region subtypes except for s2/m1 were detected. Infection with vacuolating cytotoxin producing H. pylori has been reported to be associated with particular gastro-duodenal diseases8,19,20). Several reports from Europe and the United States revealed that the positive rate of vacA s1 genotype was significantly correlated with the development of peptic ulcer diseases. These studies also showed that strains with vacA s1a allele were more likely associated with past or present ulcers than those with nonulcer dyspepsia8,19,21,22). However, our results showed that the prevalence of the s1a subtype was not statistically different between CG and either GU or DU (CG 83.3% vs. GU 64.6% or DU 76.8%). These results were substantially different from the results obtained from Western countries. Despite the overall higher prevalence of the s1a genotype, compared to the s1b genotype in ulcer diseases, the presence of the s1a genotype had no predictive value because it was found in the majority of cases, irrespective of the clinical status. These results do not support the hypothesis8) that H. pylori strain possessing specific vacA gene is useful in predicting the risk for the development of a specific clinical outcome, such as peptic ulcer diseases. Similar results were also reported by several investigators23–26). The reason for this difference on the prevalence of vacA s1 and/or the s1a alleles could be partly explained by genetic heterogeneity or structural differences in H. pylori populations throughout the world. Many studies confirmed that genomically different strains of H. pylori population appear to circulate in different geographic regions24,26,27).
The positive rate for cagA gene in our study was 94.1% (128 out of 136). Only 8 (5.9%) patients were infected with cagA-negative strains of H. pylori (5 GU, 1 DU, and 2 CG). As we previously reported15), the present study also showed that the prevalence of cagA was not different between CG and GU or DU.
Several studies especially from Europe or the United States have still shown that cagA gene is significantly correlated with the development of peptic ulcer diseases and that vacA type s1 is also strongly associated with the presence of cagA21,22,28). For example, van Doom et al.21) reported that H. pylori strains which were typed as vacA s1/cagA+/iceA could be considered the most pathogenic and were found predominantly in patients with ulcer diseases. However, there have been increasing reports that the cagA gene may not be used as a discriminator of gastroduodenal diseases in the host because the prevalence of the cagA gene is very high irrespective of the clinical status and is not related to particular gastroduodenal diseases13–16).
The overall association of the cagA positive rate with vacA s1 allele in our study was similar with the reports from others. The association was even higher in Asian strains25). However, H. pylori strains typed as the cagA+/vacA s1a were not found to be specifically associated with peptic ulcer diseases.
In conclusion, our data does not confirm the hypothesis that H. pylori strain possessing specific vacA gene is a marker to predict any specific gastroduodenal diseases in Korean patients.
Acknowledgements
The authors wish to acknowledge the financial support of the Korean Research Foundation made in the program of 1997.